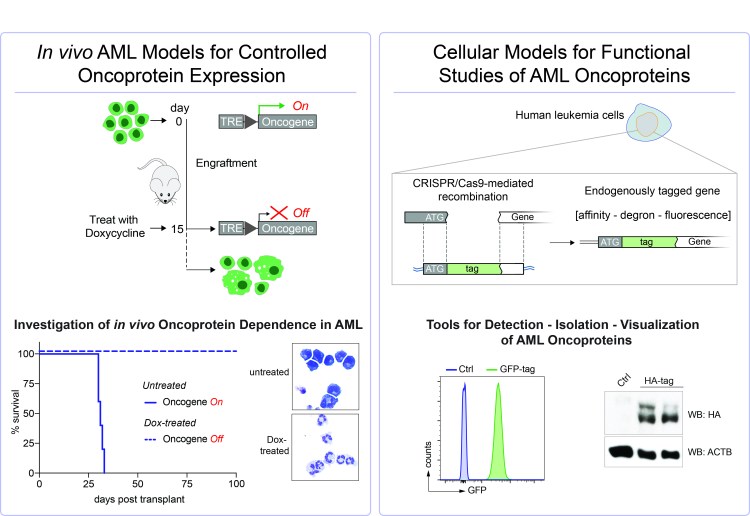
Models
We have built a comprehensive battery of tools to model and perturb oncoprotein-driven AML in cell culture and in vivo. This includes tools for inducible gene expression, modification of endogenous genes and gene silencing using RNA interference as well as CRISPR/Cas9-mediated applications.
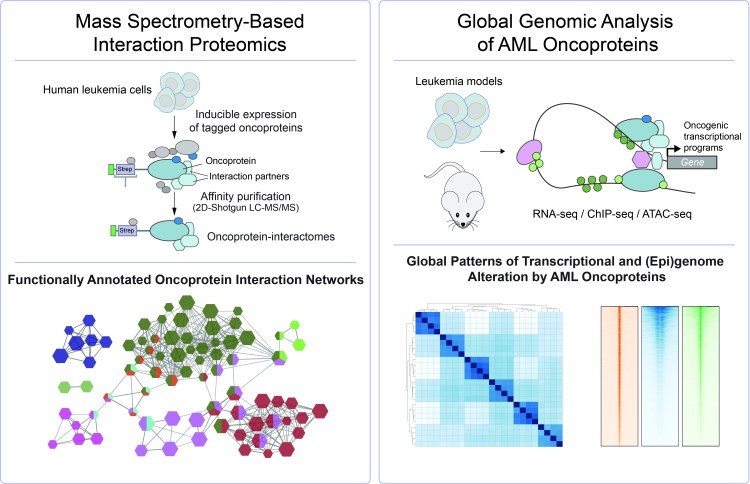
Global Approaches
We have developed a number of biochemical approaches for global proteomic characterization of AML models by Mass Spectrometry. In parallel, we use NGS-based methods to investigate global effects of AML oncoproteins on transcriptomes and epigenomes. Genome-scale CRISPR/Cas9-mediated Loss-of function screens are used to functionally explore AML genomes.
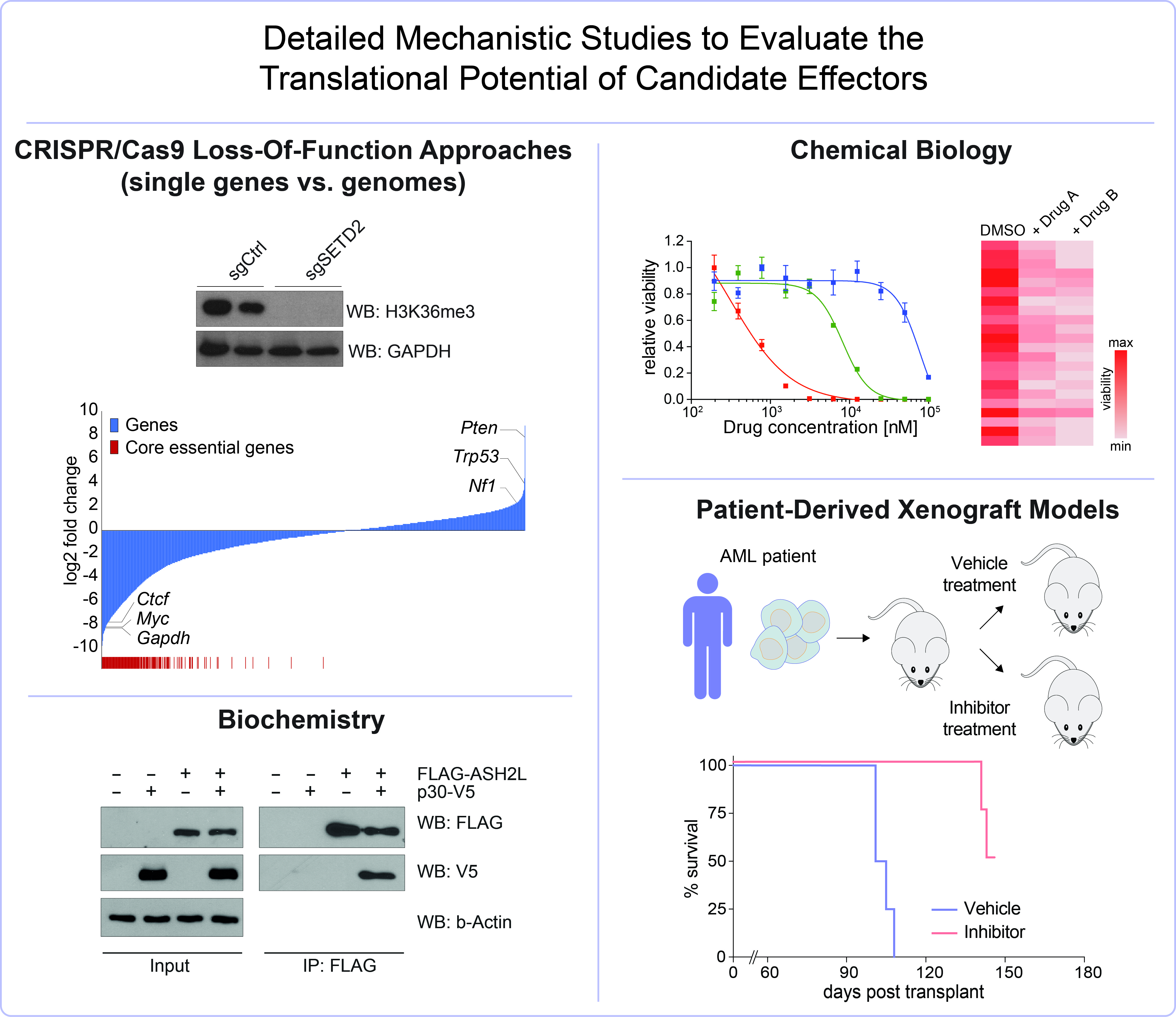
Validation
Detailed mechanistic studies are performed in AML mouse models, primary cells from AML patients and patient-derived AML xenograft models to evaluate the translational potential of our findings.